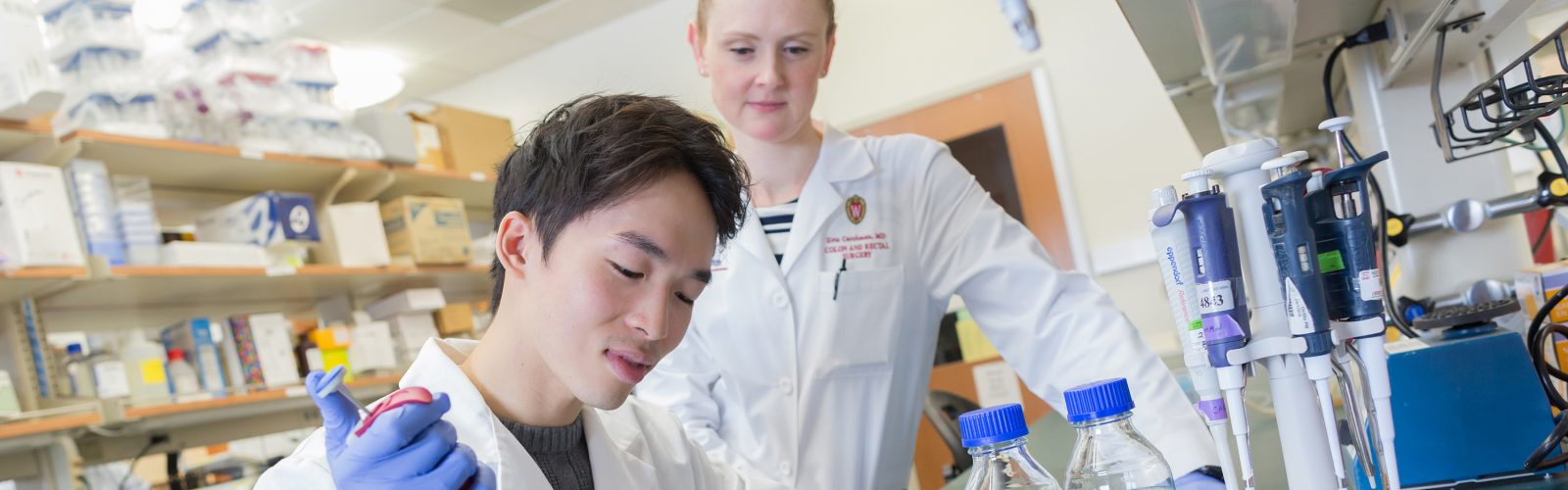
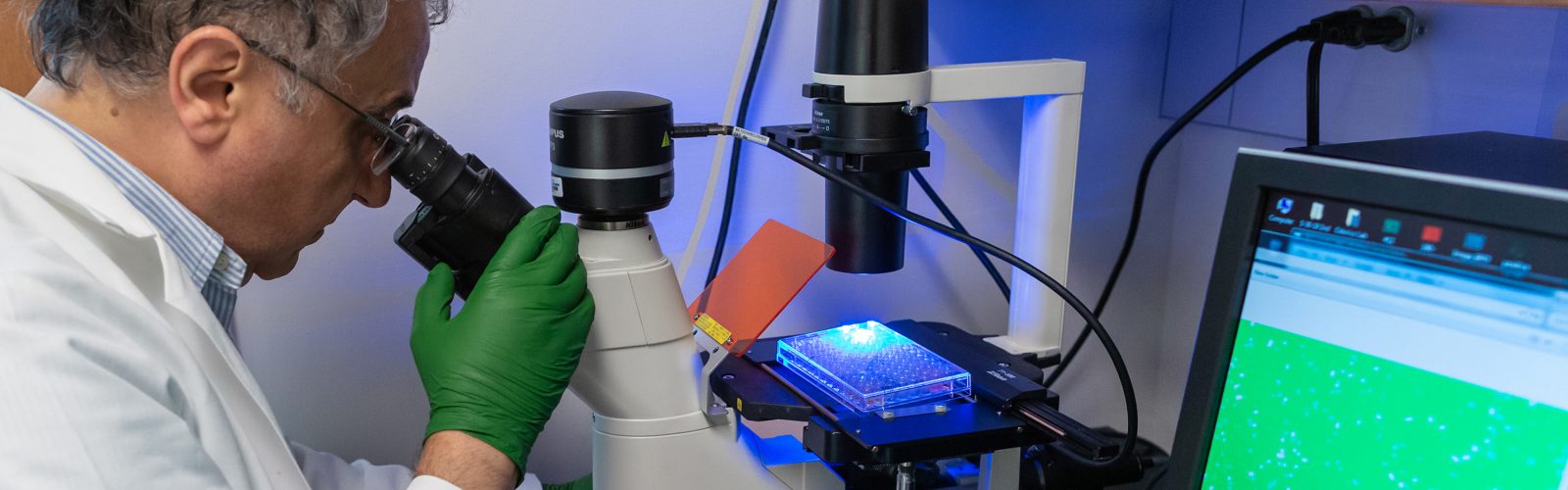
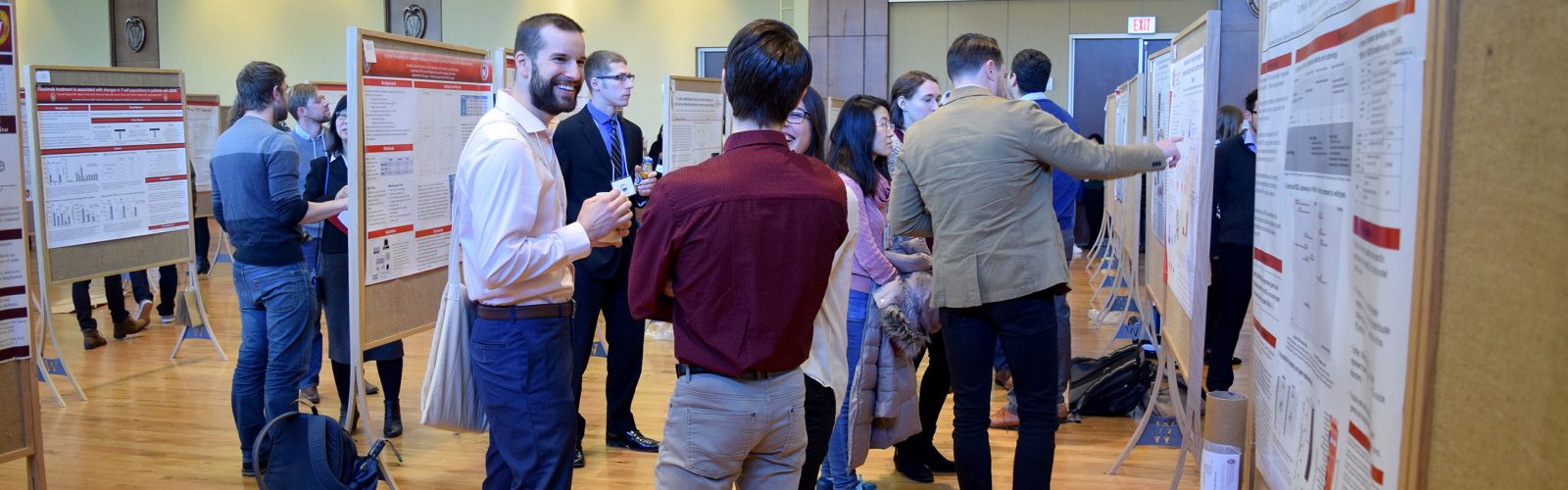
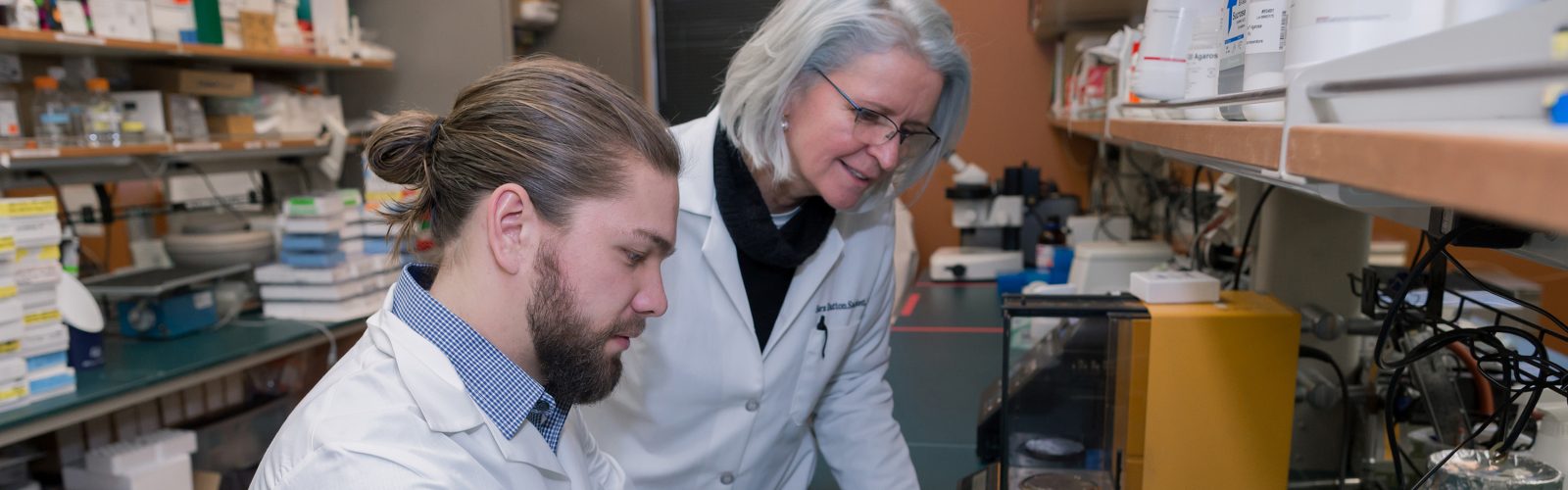
We strive to create an inclusive, collaborative and connected culture that fosters cutting-edge basic, translational, education, and health outcomes research. Our research programs investigate the underlying mechanisms of disease, translate discoveries in the laboratory to clinical trials, and offer experimental options with the goal of improving patient care. Our research funding portfolio exceeds $18 million annually with a large percentage from the National Institutes of Health.
Our Researchers & Labs >>
Latest Research News
Dr. Patrick Carney Wins Flash Talk at UW Carbone Cancer Center Research Retreat
Dr. Patrick Carney, general surgery resident, won the flash talk session at the UW Carbone Cancer Center’s Annual Research Retreat. The top six abstracts from the meeting were selected for a 10-minute oral presentation and …
Dr. Ana De Roo Awarded Prestigious Career Development Award from the American Cancer Society
Division of Colorectal Surgery Assistant Professor Ana De Roo, MD, MSc hopes to spend her career improving clinical processes and decision-making so patients with cancer are supported, informed, and receive care that is aligned with …
Opportunities for Success: Residents Thrive During Research Years
Research is one of the foundational pillars in the UW–Madison Department of Surgery. That is reflected throughout the department, including the trainee programs. All three of our residency programs provide trainees opportunities for dedicated research …
Wisconsin Surgery Research Roundup: January 2026
Wisconsin Department of Surgery members engage in remarkable research that yields many impactful publications every month. We’re highlighting several of these publications monthly to showcase the diversity of research in the department; see selections from …
Department of Surgery Celebrates 17th Annual Research Summit
On Thursday, Jan. 29, the Department of Surgery celebrated the 17th Annual Research Summit. The theme of this year’s summit was “Research Without Borders.” Our keynote speaker was Dr. Sandra Wong, Dean of Emory University …
General Surgery Resident Publishes Research about Burnout
Dr. Sydney Tan, general surgery resident, recently had her research, “Work Hours, Stress, and Burnout Among Resident Physicians,” published in JAMA Network Open. Dr. Tan and her research partners at the University of Wisconsin–Madison examined …
Odorico Lab Awarded Fall Research Competition Grant to Study How Engineered Stem Cell-Derived Islets Evade the Immune System
Over the last five years there have been significant advancements in the treatment of Type 1 diabetes, including the 2023 Food and Drug Administration approval of the first stem cell-derived therapy that can replace the …
Dr. Anna Beck Awarded Career Development Grant from UW BIRCWH Scholars Program
Congratulations are in order for Division of Surgical Oncology Assistant Professor Anna Beck, MD, who was recently awarded a position in UW’s 2025-2027 cohort of the Building Interdisciplinary Research Careers in Women’s Health (BIRCWH) scholars …
Wisconsin Surgery Research Roundup: December 2025
Wisconsin Department of Surgery members engage in remarkable research that yields many impactful publications every month. We’re highlighting several of these publications monthly to showcase the diversity of research in the department; see selections from …
Murtaza Lab Receives 2025 Badger Challenge Scholar Grant
The lab of Division of Surgical Oncology Associate Professor Dr. Muhammed Murtaza is committed to identifying and developing ways to diagnose cancer early, when more patients can be cured. Murtaza, who is also the Director …
Project led by Dr. Brigitte Smith Awarded $1.1 Million Grant from AMA
A team led by Dr. Brigitte Smith, an associate professor in the Division of Vascular Surgery and the Department of Surgery’s Vice Chair of Education, has been selected as one of 11 teams to receive …
Carchman Lab Receives Funding to Study How Precancerous Lesions Develop into Anal Cancer
Anal cancers often develop from precancerous lesions called anal dysplasia. While evidence suggests that treating these lesions can prevent the development of anal cancer, researchers still don’t understand exactly how and why a lesion changes …
Wisconsin Surgery Research Roundup: November 2025
Wisconsin Department of Surgery members engage in remarkable research that yields many impactful publications every month. We’re highlighting several of these publications monthly to showcase the diversity of research in the department; see selections from …
Surgery Scientist Receives Funding to Examine Early Immune Response in Type 1 Diabetes Transplant Therapy
Type 1 diabetes affects more than 1.6 million people in the U.S., including a growing number of children and adolescents. The disease develops when the immune system mistakenly attacks and destroys the pancreatic islet cells, …
Wisconsin Surgery Research Roundup: September 2025 and October 2025
Wisconsin Department of Surgery members engage in remarkable research that yields many impactful publications every month. We’re highlighting several of these publications monthly to showcase the diversity of research in the department; see selections from …
- More Research posts

Research Leadership

Luke Funk, MD, MPH
Vice Chair of Research

Angela Gibson, MD, PhD
Vice Chair of Research

JoAnne Vaccaro
Director of Research Operations
Clinical Trials Core
Provides coordinator and regulatory support for all types of human subjects research including sponsored clinical trials, survery studies, registries, and chart reviews.
UW Humanized Mouse Core
The Humanized Mouse Core at the University of Wisconsin–Madison provides investigators with a variety of humanized mouse models for their individual research needs.
Statistical Analysis and Research Programming (STARP) Core
Provides faculty, researchers, and residents/fellows/students with comprehensive statistical and research programming support for their research activities including study design, grant application, abstract/manuscript review, statistical analysis, programming and results interpretation.
WiSOR Qualitative Core
The WiSOR Qualitative Core offers faculty and staff consultation, support, and expertise on study design, methodology, data collection and analysis, and interpretation of results. It also offers transcription services.