You may remember learning in high school science that proteins are the building blocks of life. Now, a University of Wisconsin-Madison research team has created a huge data set on animal tissue proteins that provides building blocks for understanding how medicines work, how wounds heal, how tissue grafts change over time, and more. And they’ve made this resource available for free, along with a digital app that simplifies its use.
Department of Surgery faculty Nathan Welham, PhD, and Xudong Shi, MD, PhD, worked with colleagues from the Department of Chemistry—Lloyd Smith, PhD, Brian Frey, PhD, and (then-graduate-student) Zach Rolfs, PhD—to create a broad set of data on the rates of protein turnover in living animal tissues. Previously, a full and easily accessible data set like this did not exist, despite its relevance to biology, medicine and surgery. The team’s study was recently published in the scientific journal Nature Communications.
As Welham explains, proteins in animal tissues are very dynamic—they are constantly forming and breaking down, and even recycling into new forms. Information on how fast proteins break down, measured as their half-lives, can be extremely useful in figuring out how medicines work and how tissues heal and reform, among other body processes.
To determine protein half-lives, the researchers used a technique known as isotopic labeling. They fed three generations of mice food containing a “heavy” version of the amino acid lysine, called Lys8. They then switched the mice to food containing normal lysine (Lys0), sampled tissues from the mice at set intervals, extracted proteins from the tissue samples, and analyzed those proteins
for the amounts of Lys0 and Lys8. From that data, they were able to determine how quickly each type of protein was breaking down and being remade—in other words, the rate of protein turnover.
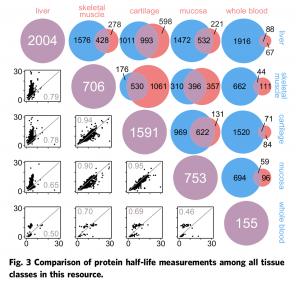
The result is a data set of the half-lives of over 3,000 distinct proteins in five different types of tissues: liver, skeletal muscle, cartilage, mucosa, and blood.
“Such a comprehensive turnover resource – generated from tissues with a wide array of functions within the body – allows us to tackle important scientific questions in ways that were previously not possible,” said Welham.
The data also served as inputs for a software program named ApplE Turnover (Application for Elucidating Protein Turnover), which team member Rolfs developed so that researchers can quickly and easily use the data set.
Dr. Frey explained, “The program allows users to search for individual proteins and then visualize their turnover rates in different tissues, along with confidence estimates and other statistical considerations.”
The team hopes their work will serve as a research asset that will ultimately lead to vital, potentially life-enhancing discoveries. The full Nature Communication article is available here, and includes information on accessing the data set and the AppIE Turnover software.